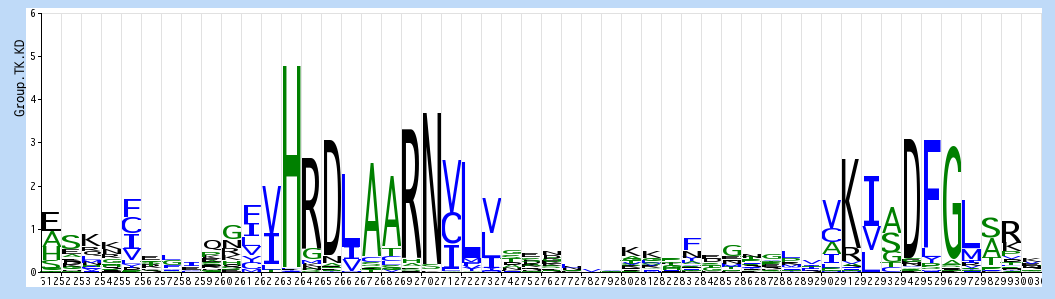

Each stack of letters is from a single position in the aligned sequences, and represents the popularity of those residues (in bits). For instance, H is almost universal at position 263, while the R at 264 is sometimes replaced by G, M or other (hard to see!) residues. If you compare this with the equivalent region in TKL group kinases:
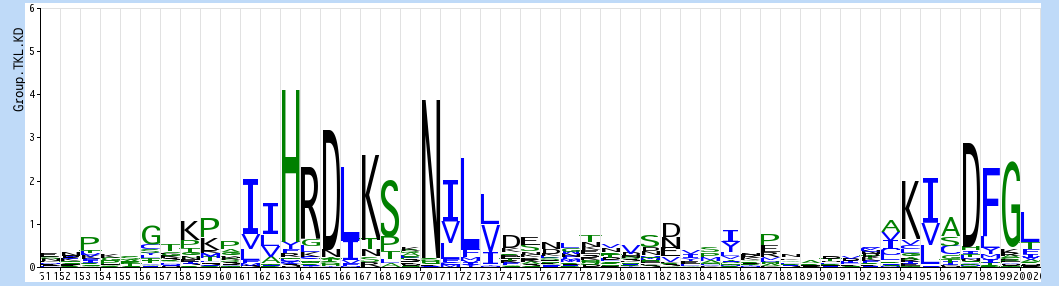
you can see the same HRD and DFG motifs standing out, but also group-specific patterns, such as the TK consensus of HRDLAARN changing to HRDLKSxN in TKL. By comparing logos, we can detect both common shared motifs and residues and motifs that are specific to individual kinase classes.
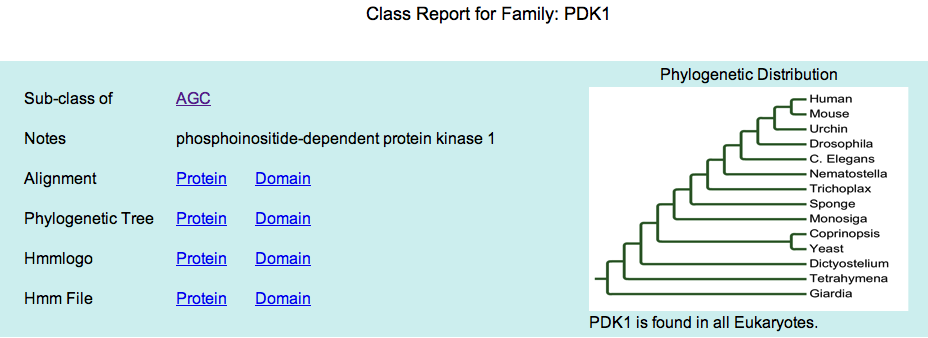
You can access both the HMM profiles and logos for every kinase class in the class report pages. Here's an example from the PDK1 family:

You can dynamically compare logos for any kinase groups, families, or subfamilies using our Logo Aligner tool, based on LogoMat (Oct 7 update: changes in HMMer and our parsing module have caused an alignment bug, which we're working on, but beware that some pairs of logos may be misaligned).